Analysis and Identification of Growth Factors found in the Amniotic Membrane, associated with Periodontal Healing.
Analysis and Identification of Growth Factors found in the Amniotic Membrane, associated with Periodontal Healing.
Gabriel Menassa1, Carole Chakar1,2, Valia Azzi3 , Robin El Jalkh1,5 , Mireille Kalassy4
1. Department of Periodontology, Faculty of Dental Medecine, Saint Joseph Univeristy of Beirut, Lebanon
2. Biology Unit, Laboratory of Cranio-Facial Research, Saint Joseph University of Beirut, Lebanon
3. Department of Ophthalmology, Bellevue Medical Center, Lebanon
4. Faculty of Science, Saint Joseph University of Beirut, Lebanon
5. Department of Digital Dentistry, Al and Evolving Technologies, Saint Joseph University of Beirut, Lebanon
*Correspondence to: Robin El Jalkh, Department of periodontology, Faculty of Dental Medicine, Saint Joseph University of Beirut, Beirut, Lebanon, Damascus Road, Beirut 1104 2020, Lebanon.
Copyright.
© 2025 Robin El Jalkh This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Received: 22 Feb 2025
Published: 03 Mar 2025
ABSTRACT
The amniotic membrane (AM) is a tissue known to be remarkably rich in growth factors. It exhibits notably anti-inflammatory and antiangiogenic effects and possesses the inherent ability to effectively induce epithelialization and cell migration at a wound site, consequently promoting efficient healing. Its pivotal role in corneal healing has been extensively studied and applied. To comprehensively explore potential applications of AM in dentistry, rigorous research was conducted on the existence and precise localization of various growth factors crucial in bone and periodontal regeneration. For this purpose, several techniques were meticulously employed, including reverse transcription-polymerase chain reaction (RT-PCR), western blot, and immunohistochemistry. The factors under scrutiny encompass Platelet-Derived Growth Factor (PDGF), Transforming Growth Factor beta (TGF-β), Insulin-like Growth Factor (IGF), and Vascular Endothelial Growth Factor (VEGF). This study findings demonstrate the presence of these different factors at both the transcriptional and translational levels, predominantly in the cytoplasm and particularly on the apical side of epithelial cells in the AM. This underscores the significant potential for the strategic exploitation of the amniotic membrane in fostering efficient bone and periodontal healing.
Keywords: Amniotic Membrane (AM), Reverse Transcription-Polymerase Chain Reaction (RT-PCR), Western Blot (WB), Immunohistochemistry (IHC), Platelet-Derived Growth Factor (PDGF), Transforming Growth Factor beta (TGF-β), Insulin-like Growth Factor (IGF), Vascular Endothelial Growth Factor (VEGF)
Analysis and Identification of Growth Factors found in the Amniotic Membrane, associated with Periodontal Healing.
Introduction
In recent years, the amniotic membrane (AM) has garnered significant attention as a compelling and readily accessible source of bioactive biological tissue enriched with growth factors and stem cells. This tissue holds immense promise for various biomedical applications, owing to its inherent regenerative properties and ease of procurement (1) . The AM represents the innermost avascular fetal membrane layer, encompassing the embryo within a sac filled with amniotic fluid. Known for its transparency, it stands as one of the thinnest human body membranes, measuring roughly 0.02-0.5 mm thick. Consisting of three identifiable layers—epithelium, basement membrane, and stromal matrix—the AM serves as a vital protective barrier during embryonic development(2).
The properties of the amniotic membrane (AM) have been shown to be multifaceted. One significant aspect is its immunomodulatory capability, aimed at regulating and maintaining a balanced immune response conducive to homeostasis (3). Various cytokines within the AM matrix play crucial roles in exerting anti-inflammatory and analgesic effects. In an experimental setup resembling an organ explant system, the human amniochorionic membrane (hACM) was exposed to lipopolysaccharide (LPS) in culture. It became evident that IL-10, present in hACM, effectively suppressed the levels of IL-6, IL-8, and TNFα (4). Other properties of the AM include anti-scarring (5), antimicrobial (6) , site-dependent angiogenic function (7,8) and tissue regeneration (9,10).
According to a recent article published in 2021(11), dermatology, orthopedics, ophthalmology, dentistry, urology, oncology, and otolaryngology were found to be the medical specialties that utilized amniotic membrane (AM) the most, particularly in the context of wound healing. Interestingly, while dermatology and orthopedics showed considerable high usage rates (32% and 26% respectively), dentistry presented with only a 6% utilization rate. Despite promising pre-clinical studies on its benefits for periodontal healing, amniotic membrane (AM) use in dentistry remains uncommon. This limited adoption can be attributed to a deficiency in comprehensive studies that specifically investigate the pertinent biological mechanisms and factors essential in dental practice.
The aim of this study was to investigate, through RT-PCR and Western blot analysis, the presence of mRNA and proteins of various growth factors that are important in dentistry, and more specifically, their localization via immunohistochemistry in the periodontal ligament, with the goal of exploring its potential in periodontal healing.
Materials and Methods
The amniotic membrane used in this study was harvested by the surgeon during a cesarean section from one healthy woman. The procedure was conducted in accordance with ethical standards and received the approval of the Ethical Committee of Saint Joseph University (Ref : USJ-2024-128). All necessary consent forms were obtained from the donor, ensuring compliance with institutional and ethical guidelines. The inclusion criteria of this study were as follow : Women aged 18-40 years , Non-smokers and no history of substance abuse , no history of obstetric complications that might affect the quality of the amniotic membrane . The exclusion criteria included positive screening for any infectious diseases, severe allergies or history of anaphylactic reactions and history of complications during current or previous pregnancies.
A. Preparation of the amniotic membrane
The placenta is harvested by the surgeon during a cesarean section. It is then transported to the tissue bank in a double sterile container, placed in a sterile bag, all within a refrigerated box.
Upon arrival at the human tissue bank, the placenta is dissected in a clean room under a vertical laminar flow hood. It undergoes washes with Ringer's lactate to remove vessels and mucus and is then incubated overnight in an antibiotic solution. After antibiotic incubation, it undergoes 6 washes with Ringer's lactate to eliminate any trace of antibiotic, then the chorion is removed and the amniotic membrane isolated.
Different segments of the membrane (2X3cm) are processed and undergo different methods of extracting mRNA and proteins as follows:
- 2 segments of membrane before detachment of the chorion are incubated in a 10% formalin solution for histological sections and immunohistochemistry.
- 2 segments of membrane without chorion are incubated in a 10% formalin solution for histological sections and immunohistochemistry.
Sample 1: 1 piece of amniotic membrane (50 cm2) is placed in RPMI (10 ml) and incubated for 4 days (37°C, 5% CO2), then the supernatant is stored at -80°C to recover the secreted factors in the medium.
Sample 2: 2 pieces of membrane are crushed in the presence of Trizol (1 ml), followed by 3 sonication cycles of 45 seconds each. The sample is then stored at -80°C for the extraction of proteins and mRNA.
Sample 3: 1 piece of membrane is crushed in the presence of 1 mL of Trizol, then stored at -80°C for the extraction of proteins and mRNA.
Sample 4: Duplicate of sample 3.
Sample 5: 2 pieces of membrane undergo a trypsin bath (500 μL) for 10 minutes at 37°C (the membrane is regularly shaken during incubation to ensure proper detachment of epithelial cells). After incubation, 40 ml of RPMI is added to dilute the trypsin effect, followed by centrifugation for 10 minutes at 4000 rpm. The pellet is then resuspended in 1 ml of Trizol and undergoes crushing followed by sonication. The sample is then stored at -80°C for the extraction of proteins and mRNA.
B. Total RNA extraction (according to the SIGMA protocol)
The RNA extraction was performed on samples 2, 3, 4, and 5.
1-Sample Treatment :
The samples in question are thawed in an ice bath, then centrifuged at 12,000 xg for 10 minutes to remove any insoluble material (membrane debris, polysaccharides...). To ensure complete dissociation of complex nucleoproteins, the samples are then incubated for 5 minutes at room temperature, followed by treatment with pure chloroform (0.2 ml/ml of Trizol used during the initial treatment of the sample). After the addition of pure chloroform, agitation for 15 seconds is necessary before allowing the samples to sit for 15 minutes at room temperature. Finally, the four tubes are centrifuged at 12,000 xg for 15 minutes at 4°C.
After centrifugation, three phases are visible in the tubes. These phases are as follows:
- The aqueous phase on the surface is transferred to a new Eppendorf tube, after which isopropanol (0.5 ml/ml of Trizol) is added. The mixture is then left for 5 to 10 minutes at room temperature after thorough mixing. Centrifugation at 12,000 xg for 10 minutes at 4°C enables the recovery of a pellet, which is subsequently washed in 75% ethanol (1 ml/ml of Trizol). Following a final centrifugation at 7500 xg for 5 minutes at 4°C, the RNA pellet is obtained. The supernatant is removed, and a quick spin is performed to recover the last drop of 75% ethanol. Subsequently, the tubes are left for 5 to 10 minutes in the air to briefly dry the pellet. Finally, the pellet is resuspended in an appropriate volume of sterile water (in our case, 20 μL), and to ensure complete dissolution of the RNA pellet, a series of 3 vortex/centrifugation cycles is performed.
-The remaining phases containing DNA and proteins are stored at -20°C for potential DNA and/or protein extractions.
2. Verification of the quality and quantity of the extracted RNA
The quality and quantity of the extracted RNA can be obtained by measuring the optical density of the samples. To achieve this, the four samples are diluted 200 times (1 μL of each sample in 199 μL of sterile water). Subsequently, the optical density is measured for each at 260 and 280 nm. Based on the optical density read at 260 nm, the calculation of the total RNA concentration is performed using the formula:
[ARN]= 40 x 200 x DO260 (Each unit of OD corresponds to 40 μg/ml)
C. Reverse Transcription-Polymerase Chain Reaction (RT-PCR)
RT-PCR facilitates the assessment of the expression of specific growth factors targeted within the amniotic membrane by amplifying particular regions of cDNA corresponding to the mRNA encoding these growth factors, provided these RNA molecules are present.
1. Selection of Appropriate Primers
The selection of primers involves multiple stages. Initially, it is imperative to identify the complete coding sequence of each protein whose expression we intend to investigate, followed by the design of suitable primers or "primer design."
Numerous possibilities arise during this process, with the selection being contingent upon two primary criteria:
- The size of the product, with preference given to larger products.
- The specificity of the product concerning the sequence we aim to amplify. To ensure this, it is essential that both chosen primers (forward & reverse) exclusively bind to our target sequence. This can be verified by performing a "blast" for each primer against the target sequence.
All these stages are facilitated by a specialized software.
The primer sequences corresponding to the selected factors for study in our project are as follows:
Table 1 : Sequences of the primers for the various growth factors (Ordered from InvitrogenÓ)
|
IGF-1 (insulin like growth factor 1):
|
|
|
PDGF-2 (platelet-derived growth factor 2):
|
PDGF-A (platelet-derived growth factor A):
|
|
VEGF (vascular endothelial growth factor)
|
TGF-beta (transforming growth factor beta)
|
2. RNA Sample Treatment with DNase: (according to the Promega protocol)
Fig 1 : RNA Sample Treatment with DNase in order to avoid RNA contamination with DNA
3. RT: (according to the Invitrogen protocol)
Before amplifying the desired regions the synthetization of complementary DNA (cDNA) from the extracted RNA was done using SuperScriptTM II RT.
The following elements are added to an Eppendorf tube:
- Random primers (0.2 µg/µL): 4 µL
- dNTP Mix (10 mM each): 4 µL
- Sterile (nuclease-free) water: 25 µL
In 3 PCR tubes (nuclease-free), the following are added:
- Treated RNA: 5 µL (equivalent to 1 µg)
- Prepared solution above: 7 µL
The samples are incubated for 5 minutes at 65 ºC, quickly cooled in an ice bath, then briefly centrifuged before adding the following to each:
- 5x First Strand Buffer: 4 µL
- 0.1M DTT: 2 µL
- RNase OUTTM: 1 µL
Finally, the samples are incubated for 2 minutes at 25 ºC before adding 1 µL of Superscript II RT followed by a series of incubations:
- 10 minutes at 25 ºC, 50 minutes at 42 ºC, 15 minutes at 70 ºC.
4. PCR
A PCR is performed for each growth factor on 2 µL of cDNA.
The mixture is prepared in an Eppendorf tube as follows:
|
10x Reaction buffer: 6 x 2.5 µL |
|
MgCl2 (50 mM): 6 x 1.5 µL |
|
dNTPs Mix (10 mM): 6 x 1 µL |
|
Left primer (10 µM): 6 x 2.5 µL |
|
Right primer (10 µM): 6 x 2.5 µL |
|
Sterile water: 6 x12.8 µL |
To this mixture, 0.2 µl Taq DNA polymerase is added before taking 23 µL and transferring them to 5 PCR tubes (each containing either a specific cDNA or a negative or positive control).
5. Revelation
To reveal the PCR results , agarose gel electrophoresis is performed. In this process, PCR products are mixed with loading dye and carefully loaded onto an agarose gel alongside a molecular weight marker. The gel is then subjected to an electric current, causing DNA fragments to migrate through the gel. Smaller fragments move faster and farther, while larger ones remain closer to the origin. After migration, the gel is visualized under UV light. The molecular weight marker helps estimate fragment sizes, negative controls ensure no contamination, while positive controls confirm successful PCR amplification.
D.Protein Extraction (according to the SIGMA protocol)
Following the initial treatment of amniotic membrane samples with pure chloroform, three phases were obtained. The first phase was collected to extract total RNA. The remaining two phases (a white phase containing DNA and an organic phase containing proteins) were frozen for potential protein extraction. It is recommended to carefully remove the remaining aqueous phase from the surface. Then, 0.3 ml/ml of absolute ethanol TriReagent is added (to precipitate DNA), mixed, and left for 2 to 3 minutes at room temperature. Centrifuging the mixture at 2000 xg for 5 minutes at 4 ºC will yield a DNA pellet and a supernatant containing proteins. The collected supernatant is combined with pure isopropanol, allowing the proteins to be collected. After leaving the tubes for 10 minutes at room temperature, they are centrifuged at 12000 xg for 10 minutes at 4 ºC. The resulting supernatant is discarded, while the protein pellet is vortexed in the presence of 2 mL of absolute ethanol and left for 20 minutes at room temperature. Centrifuging at 7500 xg for 5 minutes at 4 ºC yields a pure protein pellet, which is air-dried for 5 to 10 minutes. This pellet is then dissolved in 1% SDS (repeated pipetting) and undergoes a final centrifugation at 10,000 xg for 10 minutes at 4 ºC. The resulting supernatant from the centrifugation can then be directly used for western blotting or stored at -20 ºC. Samples 2, 3, and 5 underwent this treatment. To extract the proteins of the supernatant of the amniotic membrane (sample 1)trichloroacetic acid was added.
E. Western Blot
Gel Preparation: Polyacrylamide gel is a separation matrix used in biomolecule electrophoresis, such as proteins or DNA fragments. A concentration gel (4% stacking gel) is poured on top of the separation gel (12%) to allow homogeneous entry of samples into the separation gel. Individual lanes are created using a "comb" that divides the gel into equal portions intended for the migration of each sample.
|
Componant |
Volume |
|
ddH20 |
3,5 ml |
|
mélange d'acrylamide 30% |
4 ml |
|
1,5M Tris pH 8.8 |
2,5 ml |
|
10% SDS |
0,05 ml |
|
10% ammonium persulfate |
0,005 ml |
|
TEMED |
0,004 ml |
Table 1 : Composition of the 12% polyacrylamide separation gel
|
Componant |
Volume |
|
ddH20 |
6,1 ml |
|
mélange à 30% d'acrylamide |
1,3 ml |
|
1,0M Tris pH 6.8 |
2,5 ml |
|
10% SDS |
0,1 ml |
|
10% ammonium persulfate |
0,05 ml |
|
TEMED |
0,01 ml |
Table 2 : composition of the 4% polyacrylamide gel
After 1 hour and 30 minutes of migration at 100V, the proteins are transferred onto a PVDF membrane to come into contact with specific antibodies. The membrane is marked on the transfer surface and then transferred into 1x TBS-Tween (0.005%) + 5% skim milk. It is saturated in this buffer/milk for at least one hour at room temperature with continuous. Incubation with the first antibody corresponding to each growth factor (Table 3), diluted in TBS-Tween+ (0.5% skim milk), is performed for at least 1 hour at room temperature. A subsequent wash is done after 4 times with TBS-T (0.05%) (1x for 15 seconds and 3x for 15 minutes each). Incubation with the second antibody is carried out considering the first antibody (rabbit or mouse), in condition that it must be produced by the same species (Table 3).
The wash is then done 4 times with TBS-T (0.005%) (once quickly and three times for 15 minutes each). The antibody-peroxidase detection is performed using the ECL Amersham detection kit: the two reagents are mixed. The mixture is added to the membrane with proteins and left to incubate for 1 minute at room temperature without shaking. After removing excess reagent and wrapping the membrane in plastic wrap, a film for chemiluminescence detection is placed over the membrane in a cassette and exposed for 1 minute initially, then developed. Depending on the result, it is exposed again for different lengths of time based on the intensity of the bands.
|
|
WB |
IHC |
|||||
|
Antibodies |
Expected size after migration on SDS-PAGE gel. |
Clonality |
Dilution |
2nd anticorps correspondant (dilution) |
Dilution of the 1st antibody |
Incubation period with the 1st Ab |
Dilution of the 2nd Ab |
|
TGFß |
12 à 16 KDa 22 KDa 26 KDa 30 KDa 46 KDa |
Mouse mono clonale (ab1279) |
1/500 |
Rabbit poly clonal to mouse IgG (ab6728) (1/2000) |
1/1000 |
60 mn |
1/500 |
|
VEGF |
40 KDa |
Mouse mono- clonale (ab3109) |
1/500 |
Rabbit poly clonal to mouse IgG (ab6728) (1/2000) |
1/50 |
60 mn |
1/500 |
|
IGF |
7 KDa |
Rabbit poly clonal (ab9572) |
1/1000 |
Goat poly clonal to rabbit IgG (ab6721) (1/3000) |
1/200 |
30 mn |
1/500 |
|
PDGF |
27 KDa |
Rabbit poly clonal (ab15499) |
1/200 |
Goat poly clonal to rabbit IgG (ab6721) (1/3000) |
1/100 |
30 mn |
1/500 |
Table 3 : Specifications of antibodies used in Western blotting (WB) and immunohistochemistry (IHC).
G. Immunohistochemistry
To locate growth factors in the amniotic membrane, we conducted immunohistochemical analysis on paraffin-embedded tissue. The process involved tissue fixation in 10% formalin, followed by paraffin inclusion. Sections of 4 μm thickness were made, either containing the membrane+chorion or just the membrane. After deparaffinization and hydration, antigenic sites were unmasked using microwave treatment in citrate buffer. Subsequent steps included primary antibody incubation, followed by washes and secondary antibody incubation. The revelation was achieved using DAB solution, followed by washing and hematoxylin incubation.
Results
A-RT-PCT
Before initiating PCR amplification of the desired cDNA sequences, it is essential to verify the outcome of the reverse transcription process to ensure the presence and quality of the cDNA samples. To achieve this, the initial PCR focused on the gene encoding β-actin, a protein expressed by all cells, serving as an indicator of successful reverse transcription if cDNA is present.
1. β-actine PCR :
The reverse transcriptions conducted both yielded a positive result for amplifying β-actin. A band of the expected size (142 bp) was observed on a 2.5% agarose gel. In lane 3, the observed band acted as a positive control, confirming the successful PCR. The presence of cDNA in samples 3 and 5 was confirmed, albeit in lower quantities. No band was observed in sample 2, suggesting potential mRNA degradation due to mishandling. This genetic material was used for subsequent PCR series.
Additionally, the β-actin PCR helps normalize band intensities, aiding in determining relative quantities of PCR products for the investigated growth factors. It's noteworthy that the observed bands were of low intensity, indicating limited cDNA production.
Figure 2: PCR results of the gene encoding β actin. The well contents from left to right are as follows: well 1: size marker, well 2: negative control, well 3: positive control (PCR performed on genomic DNA), wells 4, 5, 6: PCR product from samples 2, 3, 5. The size of the PCR product is 142 bp.
2. IGF-1 PCR :
Figure 3: PCR results of the cDNA of the gene encoding IGF : the well contents from left to right are as follows: well 1: size marker, well 2: negative control, well 3: positive control (PCR performed on genomic DNA), wells 4, 5, 6: PCR product from samples 2, 3, 5. The size of the PCR product is 242 bp.
The IGF PCR demonstrated that the IGF’s mRNA is present in samples 3 and 5. Therefore , the expression of IGF at the transcriptional level in amniotic cells is confirmed.
3.PDGF-A PCR :
Figure 4: PCR results of cDNA from the PDGF-A gene. The well contents from left to right are as follows: well 1: size marker, well 2: negative control, well 3: positive control (PCR performed on genomic DNA), wells 4, 5, 6: PCR product from samples 2, 3, 5. The size of the PCR product is 577 bp.
The PCR analysis of PDGF-A revealed a single band at the expected size of 577 bp, detected only in sample 3. This confirms the presence of PDGFA mRNA and thus its expression in the amniotic membrane.
4. PDGF-B PCR :
Figure 5: PCR results of the cDNA of the PDGF-B gene. The well contents are as follows from left to right: well 1: size marker, well 2: negative control, well 3: positive control (PCR performed on genomic DNA), wells 4, 5, 6: PCR product from samples 2, 3, 5. The size of the PCR product is 169 bp.
The PCR results for PDGF-B did not reveal any bands in their amplification profile. With the positive control displaying a band at the expected size of 169 bp, it can be confirmed that PDGF-B is not expressed in the amniotic membrane.
5. VEGF PCR :
Figure 6: PCR results of cDNA from the gene encoding VEGF. The contents of the wells are as follows from left to right: well 1: size marker, well 2: negative control, well 3: positive control (PCR performed on genomic DNA), wells 4, 5, 6: PCR product from samples 2, 3, 5. The size of the PCR product is 226 bp.
VEGF mRNA is detected in both samples 3 and 5, with bands appearing at the expected size of 226 bp, confirming its expression in the amniotic tissue. It's worth noting that the band in the positive control appears larger, likely due to the primers targeting different exons.
B-Western Blot
After analyzing the growth factors' expression at the transcriptional level, their presence is studied in the membrane at the translational level. This involves two experimental conditions: searching for secreted proteins (in the supernatant, samples 1 and 1') and those remaining in the amniotic membrane (samples 2, 3, 5). To achieve this, proteins are extracted from the same samples used for RNA extraction (samples 2, 3, 5), while proteins from the medium where the membrane was incubated for 4 days are recovered by precipitation with trichloroacetic acid (Sample 1 and its duplicate 1'). Subsequently, different protein extracts are migrated on SDS gels and transferred to PVDF membranes, followed by incubations with various antibodies.
Figure 7: Membrane onto which different protein extracts labeled with anti-IGF antibody were transferred. The expected molecular weight is 7 kDa. The content of the wells is as follows from right to left: well 1, molecular weight marker; well 2, supernatant extract; well 3, supernatant extract; wells 4, 5, 6, proteins extracted from samples 2, 3, 5.
The WB results, obtained after hybridization with the anti-IGF antibody, revealed a distinct band at the expected size of 7 kDa. This band was detected both in the amniotic fluid (Figure 8, samples 1 and 1') and in well 5. These findings indicate that IGF is predominantly secreted, with a substantial amount also found in the tissue.
Figure 8: Membrane on which various protein extracts labeled with the rabbit polyclonal anti PDGF-BB antibody were transferred. The expected molecular weight is 27 kDa. The content of the wells is as follows from left to right: well 1, molecular weight marker; well 2, supernatant extract; well 3, supernatant extract; wells 4, 5, 6, proteins extracted from samples 2, 3, 5.
Considering that the TGF-β growth factor has various isoforms, it can be detected at 12, 16, 22, 26, 30, and 46 kDa. In our study, the membrane labeling with the anti-TGF-β antibody revealed bands at 16 kDa present in all samples and another at 12 kDa specifically in samples 1, 1', and 5.
Figure 9: Membrane onto which different protein extracts labeled with rabbit polyclonal anti-VEGF antibody were transferred. The expected size is 19 to 22 kDa. The contents of the wells are as follows from left to right: well 1 molecular weight marker, well 2 supernatant extract, well 3 supernatant extract, well 4, 5, 6 proteins extracted from samples 2, 3, 5.
The anti-VEGF antibody labeling reveals bands at 30 kDa in samples 1, 1’ to 6, a band at 19 kDa in sample 3, and another at 38 kDa in samples 3 and 5. According to different sources, VEGF can vary in molecular weight from 19 to 40 kDa depending on the cell source.
C. Immunohistochemistry
Following the confirmation of key growth factors associated with periodontal healing in the amniotic membrane (MA) at both the RNA and protein levels, a more precise localization was pursued through immunohistochemistry. This step helps in identifying the exact locations of these factors, particularly if they were not secreted, on histological sections.
Figure 10: Sections illustrating the presence of various growth factors in the amniotic membrane. All sections are captured at 100x magnification.
Please click here to view all images
a- The histological section is incubated for 60 minutes with the first anti-VEGF antibody.
b- The histological section is incubated for 30 minutes with the first anti-IGF antibody.
c- The histological section is incubated for 30 minutes with the first anti-TGF antibody.
d- The histological section is incubated for 30 minutes with the first anti-PDGF antibody.
Discussion
The growth factors we examined are pivotal in periodontal healing, yet their expression in the amniotic membrane (AM) remains largely unexplored in the literature. Employing a combination of RT-PCR, western blot analysis, and immunohistochemistry, our study successfully localized these factors within the AM.
These findings offer crucial insights that could significantly advance the potential application of the amniotic membrane in dental surgery. For instance, IGF-1 induction has been demonstrated in cells cultured on the AM, consistent with its transcriptional presence in the AM (Lee JH, 2006). Furthermore, we observed its predominant presence in the membrane's supernatant and epithelial cell cytoplasm at the protein level.
Regarding VEGF, while its expression in the placenta, particularly the amnion, has been established, our study adds to the understanding by detecting its mRNA presence and confirming protein expression in epithelial cell cytoplasm (Kawano Y, 2005). However, the specificity of labeling in the membrane's supernatant requires further confirmation.
PDGF, despite its critical role in periodontal regeneration, has not been thoroughly studied in the AM. Our detection of PDGF-A transcriptional expression and confirmation of its presence in epithelial cells through immunohistochemistry shed light on its potential role in periodontal healing.
Our study also confirms the existence of TGFβ in the AM, consistent with previous research implicating it in periodontal regeneration. This molecular study provides a foundational reference for the clinical development of the AM in dentistry, particularly in periodontal regeneration.
Moving forward, complementing this research with clinical trials can further explore the rich potential of the AM in various growth factors' expression and its interaction with the periodontal environment. This may lead to innovative applications, such as using the AM as a graft to stimulate bone regeneration and wound healing in dental surgery or as a patch to promote periodontal and bone regeneration.
However, before clinical application, thorough studies, including animal trials, are necessary to understand how AM growth factors promote bone growth through various mechanisms such as osteoconduction, osteopromotion, osteoinduction, and osteoformation.
The AM's expression of both angiogenic and gingival growth factors underscores its versatility in periodontal regeneration. Further investigation into its potential, particularly in bone regeneration, is warranted, given the presence of multipotent stem cells like BM-MSCs within the AM (Kim, et al., 2007).
Overall, these results present promising avenues for leveraging the AM in periodontology, akin to its remarkable success in ophthalmic surgery. Its unique capabilities and potential for improvement highlight its significance in both therapeutic and surgical domains, offering exciting possibilities for future research and application.
Conflict of interests: The authors declare no conflict of interest relevant to the content of this article.
Funding information: No funding was received to assist with the preparation of this manuscript, and the authors have no known competing financial interests.
References
1. Etchebarne M, Fricain JC, Kerdjoudj H, Di Pietro R, Wolbank S, Gindraux F, et al. Use of Amniotic Membrane and Its Derived Products for Bone Regeneration: A Systematic Review. Front Bioeng Biotechnol [Internet]. 2021 May 11 [cited 2024 Mar 16];9.
2. Law EJ, Taib H, Berahim Z, Law E, Taib H, Berahim Z. Amniotic Membrane: An Approach to Periodontal Regeneration. Cureus [Internet]. 2022 Aug 9 [cited 2024 Mar 16];14(8).
3. Sehar I, Kaul A, Bani S, Pal HC, Saxena AK. Immune up regulatory response of a non-caloric natural sweetener, stevioside. Chem Biol Interact. 2008 May 28;173(2):115–21.
4. Munoz-Torres JR, Martínez-González SB, Lozano-Luján AD, Martínez-Vázquez MC, Velasco-Elizondo P, Garza-Veloz I, et al. Biological properties and surgical applications of the human amniotic membrane. Front Bioeng Biotechnol [Internet]. 2023 Jan 9 [cited 2024 Mar 17];10.
5. Zhu YT, Li F, Zhang Y, Chen SY, Tighe S, Lin SY, et al. HC-HA/PTX3 Purified From Human Amniotic Membrane Reverts Human Corneal Fibroblasts and Myofibroblasts to Keratocytes by Activating BMP Signaling. Invest Ophthalmol Vis Sci. 2020 May 11;61(5):62.
6. Mao Y, Hoffman T, Singh-Varma A, Duan-Arnold Y, Moorman M, Danilkovitch A, et al. Antimicrobial Peptides Secreted From Human Cryopreserved Viable Amniotic Membrane Contribute to its Antibacterial Activity. Sci Rep. 2017 Oct 20;7(1):13722.
7. Küçükerdönmez C, Akova YA, Altinörs DD. Vascularization is more delayed in amniotic membrane graft than conjunctival autograft after pterygium excision. Am J Ophthalmol. 2007 Feb;143(2):245–9.
8.Human Amniotic Membrane and Its Anti-cancer Mechanism: a Good Hope for Cancer Therapy | SN Comprehensive Clinical Medicine . [cited 2024 Mar 17]. Available from: https://link.springer.com/article/10.1007/s42399-019-00090-5
9. Toda A, Okabe M, Yoshida T, Nikaido T. The potential of amniotic membrane/amnion-derived cells for regeneration of various tissues. J Pharmacol Sci. 2007 Nov;105(3):215–28.
10. Properties and Therapeutic Potential of Human Amniotic Membrane [Internet]. [cited 2024 Mar 17]. Available from: https://scialert.net/abstract/?doi=ajd.2015.1.12
11. Nejad AR, Hamidieh AA, Amirkhani MA, Sisakht MM. Update review on five top clinical applications of human amniotic membrane in regenerative medicine. Placenta. 2021 Jan 1;103:104–19.
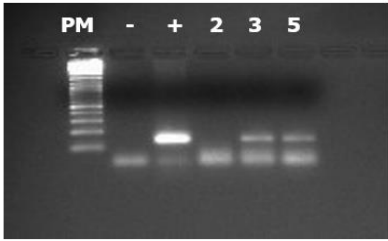
Figure 1
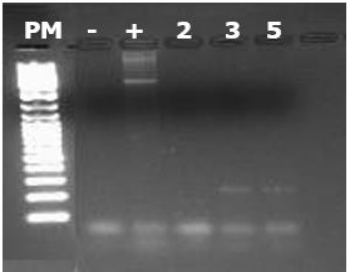
Figure 2
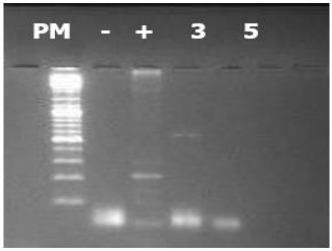
Figure 3
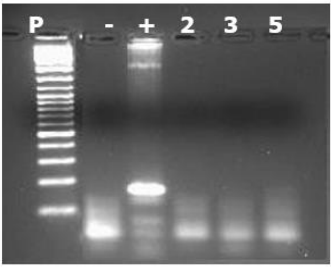
Figure 4
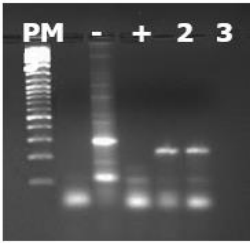
Figure 5
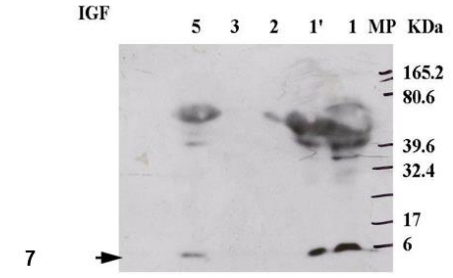
Figure 6
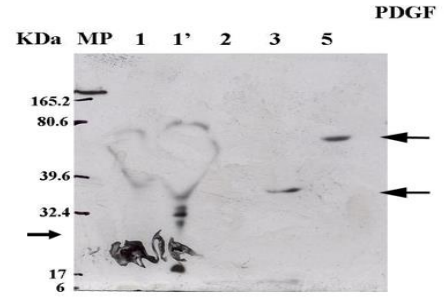
Figure 7
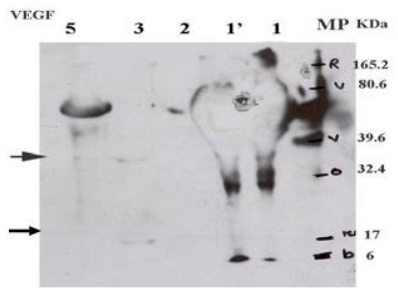
Figure 8
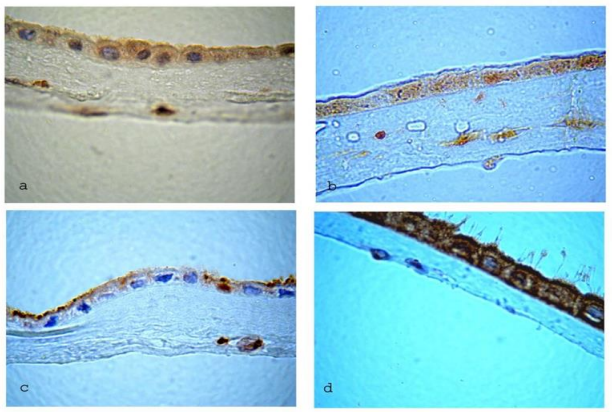
Figure 9
